Targeted Protein Degradation
Targeted Protein Degradation (TPD) harnesses natural biological processes to completely remove a disease-causing protein from the cell. This is different from traditional drugs, which typically work by binding to a disease-causing protein to correct its behavior. Degradation has a number of important benefits, including enhanced activity, reduced dose, and the ability to target undruggable proteins.
Degraders are shifting drug development paradigms from focusing on inhibition to removing pathogenic proteins.
We are focused on using AI approaches to unlock the existing prospective design limitations of molecular glue degraders, specifically to enable drugging the most important disease-causing proteins that are currently considered undruggable.
Molecular glue degraders can repurpose the cell’s natural protein homeostatic control processes to target disease-causing proteins. Our current focus is specifically on the ubiquitin-proteasome system (UPS), which has been clinically validated with molecular glue degrader drugs. Still, it has a significant need for disruptive technologies to unlock the full potential of this field.
The UPS is present in all eukaryotic cells and selectively removes proteins in three steps:
- The ubiquitin-activating enzyme (E1) activates ubiquitin, which is then transferred to a ubiquitin-conjugating enzyme (E2).
- Ubiquitin on the E2 enzyme is conjugated to lysine residues within the protein of interest, or the N-terminal amino group, by a ubiquitin (E3) ligase. E3 ligases can be highly selective due to protein-protein interactions between the E3 and the protein of interest.
- Additional ubiquitin molecules are conjugated onto the first, forming a polyubiquitinated adduct recognized by the proteasome leading to degradation of the protein of interest.
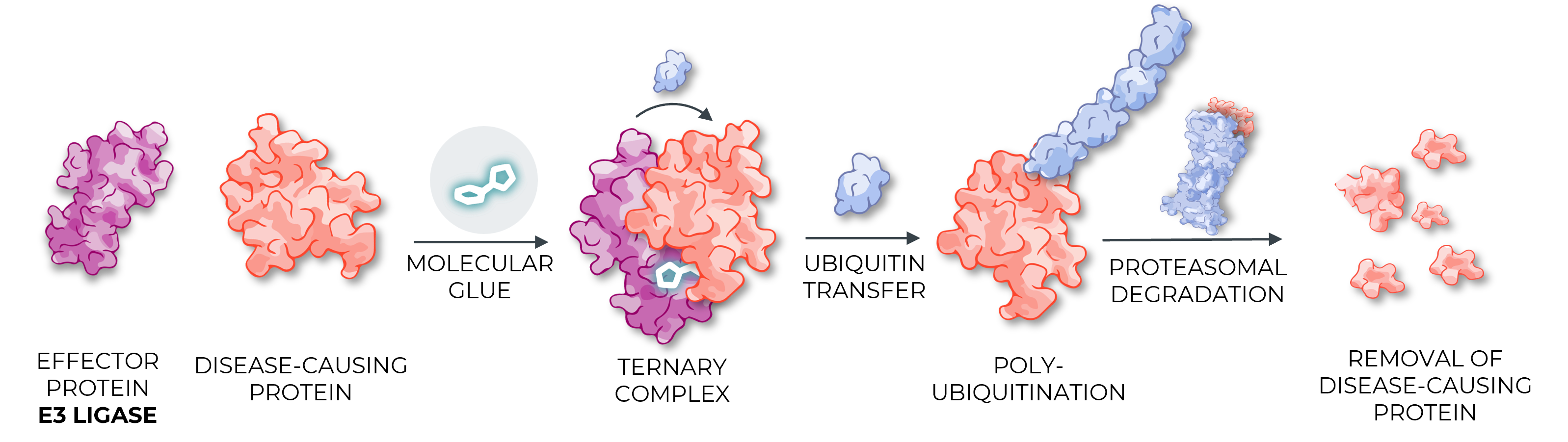
Molecular Glues
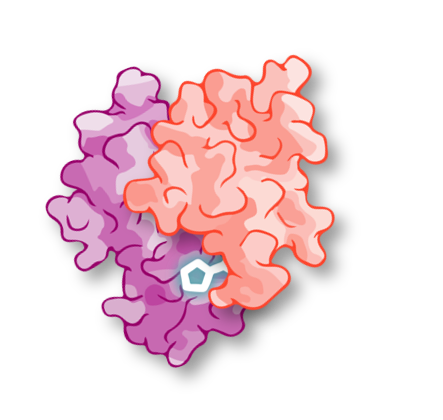
Molecular glues are next-generation degrader small molecules that stabilise interactions between the disease-causing protein and an E3 ligase. Unlike heterobifunctional degraders (such as PROTACs), which suffer from developability challenges, molecular glues have properties similar to those of traditional small-molecule drugs, enabling oral administration and CNS penetration. Marketed drugs that act as molecular glues include Revlimid, Rapamycin, and Tacrolimus.
The discovery of molecular glues is significantly limited by a lack of technologies that enable their prospective design. Current approaches require expensive and time-consuming experimentation, such as high throughput screening and extensive biophysical characterisation.
Area and Disease Agnostic
Our platform is designed with versatility at its core, transcending traditional therapeutic boundaries to address a wide spectrum of diseases. Whether it’s targeting central nervous system disorders, cancer, or other complex conditions, our molecular glue technology and AI-driven design capabilities are area agnostic. This adaptability ensures that our differentiated approach can be applied universally, offering groundbreaking treatments across various medical fields and transforming the landscape of modern medicine.
Advantages
Target Intractable Proteins
Only 10-20% of all pathogenic proteins have well-defined binding sites that can be targeted with traditional small molecule drugs. Molecular Glues do not require these well-defined binding sites and can degrade otherwise undruggable proteins. This gives promise in overcoming yet-to-be-drugged targets connected to diseases with high unmet medical needs, such as Parkinson’s. Many ongoing clinical studies back these claims. 1, 2, 3, 4, 5
High Target Selectivity via Protein-Protein Interactions
Due to their characteristic of stabilizing cellular machinery and target protein interactions, Molecular Glues can exhibit differentiated and improved selectivity compared to traditional small molecule drugs. 8, 9, 10
Ease of Manufacturing
Molecular Glues are chemically-derived small molecules and not extracted from living organisms. As a result, chemical synthesis can be scaled efficiently once the synthesis pathway is identified.
Improved Safety
The intracellular degradation machinery can vary between tissues and cells. This provides the opportunity to direct the drug to the desired site of action, reducing side effects related to activity in undesired tissues. 7, 12, 13
Disrupting Scaffolding Functions
Although small-molecule inhibitors silence distinct domains of proteins, the scaffolding functions of targeted proteins outside the small-molecule binding site can remain active. By regulating the overall protein concentration, molecular glue degraders are more likely to recapitulate the phenotypic data seen in knockout (CRISPR) and knockdown (RNAi) target validation assays.
Oral
Bioavailability
Therapeutic strategies which modulate gene function by blocking gene expression might have a similar impact on phenotypes compared to a PIC™ strategy. However, knockout (CRISPR) and knockdown (RNAi) therapies have significant administration limitations, whereas molecular glues provide the unique pharmacodynamic advantages of small molecules – including oral bioavailability.
Potency Disconnected from Affinity
Inhibitors rely on the occupancy of a binding site, which requires high affinity or high concentration to compete with endogenous substrates. Due to their event-driven pharmacology, molecular glues are catalytic in nature and consequently show higher potency. 12, 13, 14
Clinical Validation
While other molecular glue-based modalities are being explored, our focus is on molecular glue degrades that harness the ubiquitin-proteasome system to induce targeted protein degradation. Molecular glue degrades have already been approved in the markets.
-
Kathleen M. Sakamoto, Kyung B. Kim, Akiko Kumagai, Frank Mercurio, et al., (2001). Protacs: Chimeric molecules that target proteins to the Skp1–Cullin–F box complex for ubiquitination and degradation. PNAS 15, 8554-8559
-
Asher Mullard (2021). Targeted protein degraders crowd into the clinic. News, Nature Reviews Drug Discovery 20, 247-250.
-
Eric S. Wang, Alyssa L. Verano, Radosław P. Nowak, et al., (2021).
Acute pharmacological degradation of Helios destabilizes regulatory T cells Nature Chemical Biology. 17, 711–717 -
Schneider, M., Radoux, C.J., Hercules, A. et al. The PROTACtable genome. Nat Rev Drug Discov (2021). https://doi.org/10.1038/s41573-021-00245-x
-
Pujols J, Peña-Díaz S, Lázaro DF, Peccati F, Pinheiro F, et al., (2018). Small molecule inhibits α-synuclein aggregation, disrupts amyloid fibrils, and prevents degeneration of dopaminergic neurons. PNAS 41, 10481-10486
-
Piazza I, Beaton N, Bruderer R, Knobloch T, Barbisan C, Chandat L, et al., (2020). A machine learning-based chemoproteomic approach to identify drug targets and binding sites in complex proteomes. Nature communications 11, 4200. doi:10.1038/s41467
-
Kostic, M., & Jones, L. H. (2020). Critical Assessment of Targeted Protein Degradation as a Research Tool and Pharmacological Modality. Trends in pharmacological sciences, 41(5), 305–317.
-
Smith, B.E., Wang, S.L., Jaime-Figueroa, S. et al. Differential PROTAC substrate specificity dictated by orientation of recruited E3 ligase. Nat Commun 10, 131 (2019).
-
Bondeson DP, Smith BE, Burslem GM, Buhimschi AD, Hines J, Jaime-Figueroa S, Wang J, Hamman BD, Ishchenko A, Crews CM. Lessons in PROTAC Design from Selective Degradation with a Promiscuous Warhead. Cell Chem Biol. 2018 Jan 18;25(1):78-87.e5. doi: 10.1016/j.chembiol.2017.09.010. Epub 2017 Nov 9. PMID: 29129718; PMCID: PMC5777153.
-
Philipp M. Cromm, Craig M. Crews,Targeted Protein Degradation: from Chemical Biology to Drug Discovery (2017), Cell Chemical Biology, (24)1181-1190,
-
Lai AC, Crews CM. Induced protein degradation: an emerging drug discovery paradigm. Nat Rev Drug Discov. 2017;16(2):101-114. doi:10.1038/nrd.2016.211
-
Guo, WH., Qi, X., Yu, X. et al. Enhancing intracellular accumulation and target engagement of PROTACs with reversible covalent chemistry. Nat Commun 11, 4268 (2020).
-
Proof-of-Concept with PROTACs in Prostate Cancer (2020) Cancer Discov. (10) (8) 1084
-
Katherine A. Donovan, Fleur M. Ferguson, Jonathan W. Bushman et al., (2020). Mapping the Degradable Kinome Provides a Resource for Expedited Degrader Development. Cell. 183, 1714-1731